# ==============================================================================
# SECTION D: OPTIMIZED SAMPLING BIAS ANALYSIS (STUDY REGION LIMITED)
# ==============================================================================
cat("\n", paste(rep("=", 70), collapse = ""), "\n")
cat(" SECTION D: OPTIMIZED SAMPLING BIAS DETECTION\n")
cat(paste(rep("=", 70), collapse = ""), "\n")
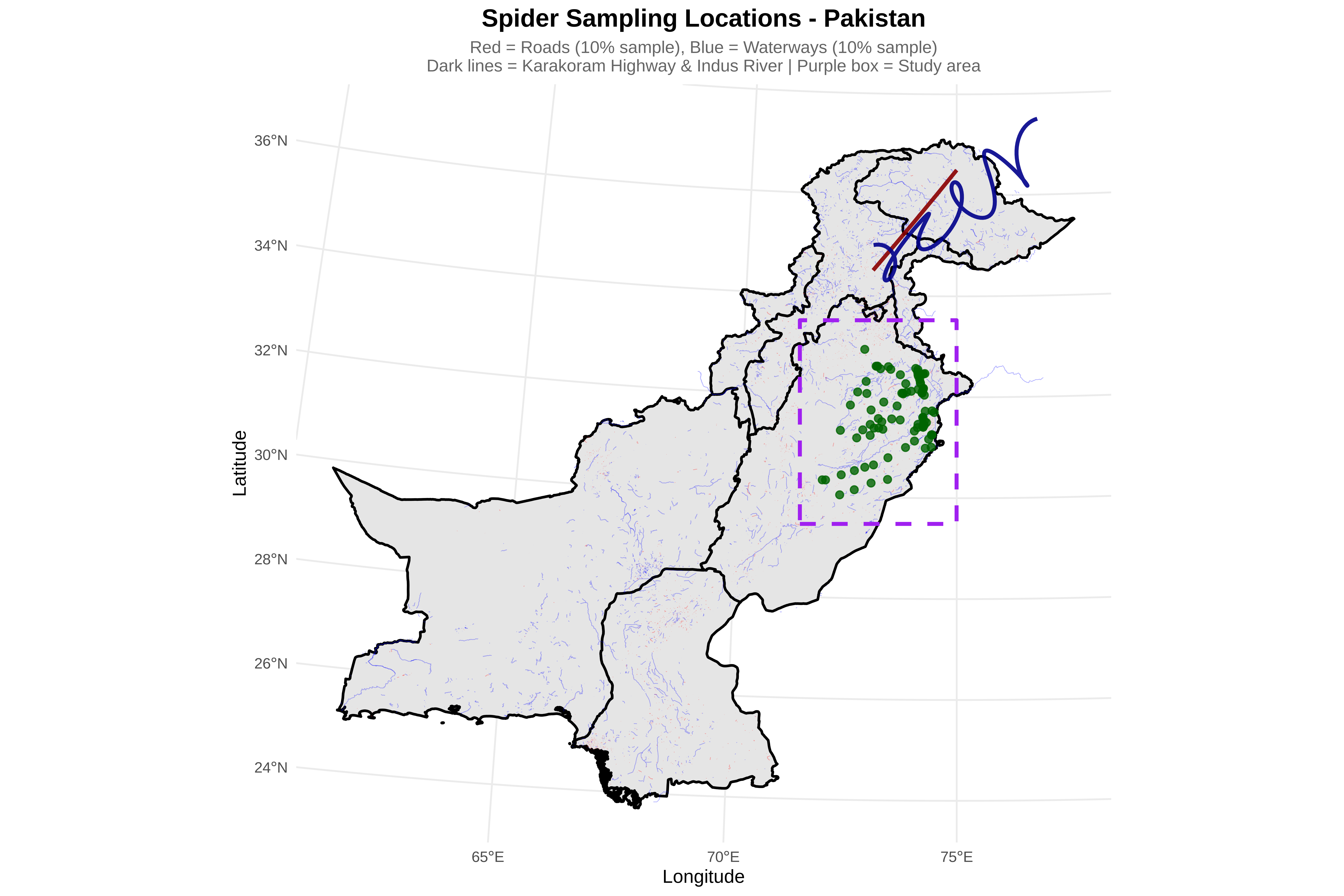
# D1: Define study region (buffer around spider points)
cat("\n1. Defining study region...\n")
# Create buffer coordinates (50 km buffer)
study_buffer <- 50000 # 50 km in meters
xmin_buffer <- spider_bbox$xmin - study_buffer
xmax_buffer <- spider_bbox$xmax + study_buffer
ymin_buffer <- spider_bbox$ymin - study_buffer
ymax_buffer <- spider_bbox$ymax + study_buffer
cat("Buffer coordinates:\n")
cat(" xmin:", xmin_buffer, "\n")
cat(" xmax:", xmax_buffer, "\n")
cat(" ymin:", ymin_buffer, "\n")
cat(" ymax:", ymax_buffer, "\n")
# Create study region polygon
study_polygon <- st_polygon(list(matrix(c(
xmin_buffer, ymin_buffer,
xmax_buffer, ymin_buffer,
xmax_buffer, ymax_buffer,
xmin_buffer, ymax_buffer,
xmin_buffer, ymin_buffer
), ncol = 2, byrow = TRUE)))
study_region_sfc <- st_sfc(study_polygon, crs = st_crs(spiders_utm))
cat("Study region size:", round((xmax_buffer - xmin_buffer)/1000, 1), "km x",
round((ymax_buffer - ymin_buffer)/1000, 1), "km\n")
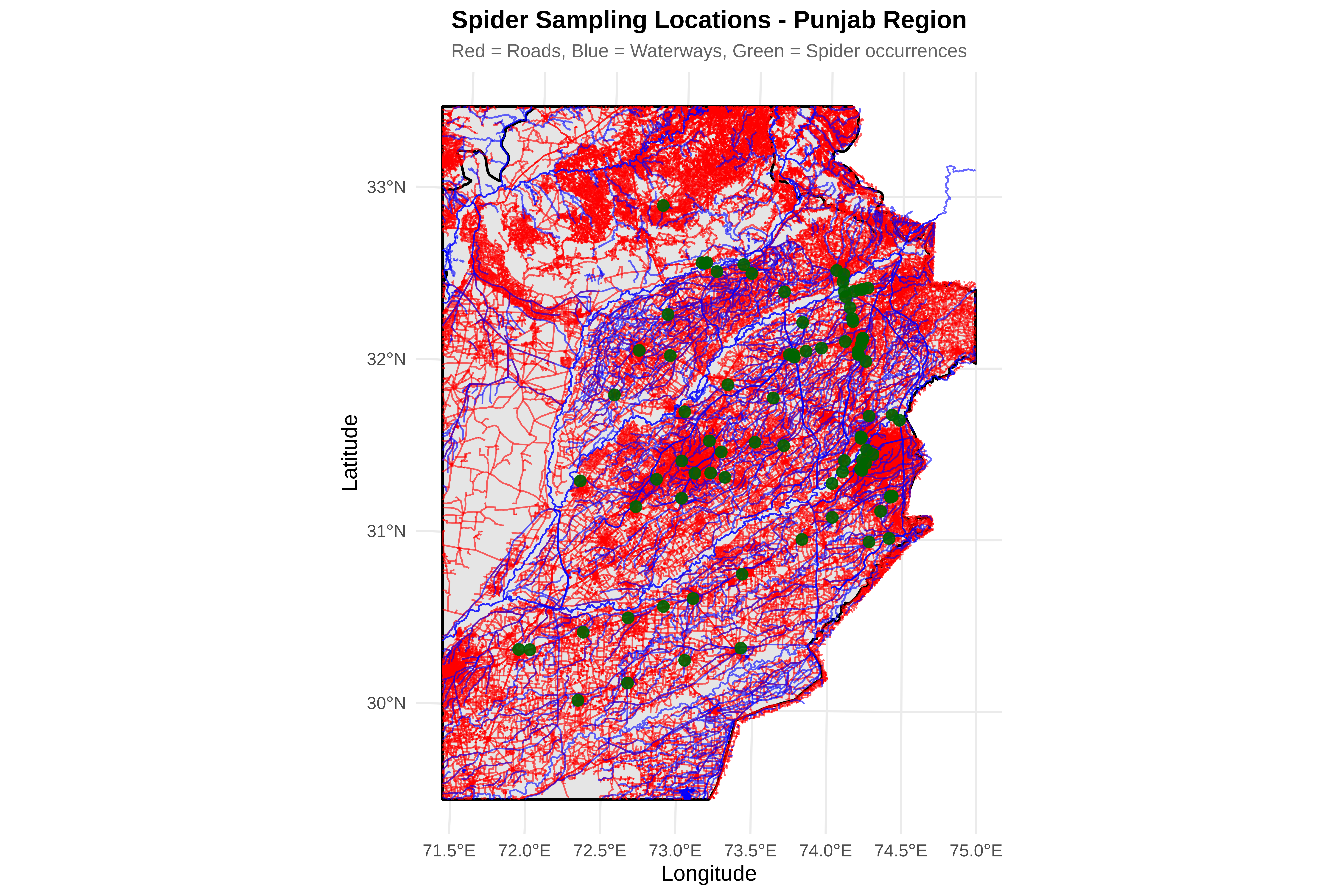
# D2: Crop roads and rivers to study region
cat("\n2. Cropping features to study region...\n")
# Function to crop and validate
crop_features <- function(features, region, feature_name) {
cat(paste(" Cropping", feature_name, "..."))
indices <- st_intersects(region, features)[[1]]
if(length(indices) > 0) {
cropped <- features[indices, ]
cat(" kept", nrow(cropped), "features (",
round(100 * nrow(cropped) / nrow(features), 1), "% of total)\n")
return(cropped)
} else {
cat(" WARNING: No features found!\n")
return(features[0,])
}
}
roads_cropped <- crop_features(roads_utm, study_region_sfc, "roads")
rivers_cropped <- crop_features(rivers_utm, study_region_sfc, "rivers")
# D3: Extract coordinates for nearest neighbor search
cat("\n3. Extracting coordinates for nearest neighbor search...\n")
extract_coords <- function(sf_object) {
coords <- st_coordinates(sf_object)
return(coords[, 1:2, drop = FALSE])
}
cat(" Extracting road coordinates...")
road_coords <- extract_coords(roads_cropped)
cat(" done (", nrow(road_coords), "points)\n")
cat(" Extracting river coordinates...")
river_coords <- extract_coords(rivers_cropped)
cat(" done (", nrow(river_coords), "points)\n")
# D4: Calculate distances for presence points
cat("\n4. Calculating distances for presence points...\n")
presence_coords <- st_coordinates(spiders_utm)
cat(" Finding nearest roads...")
road_nn <- get.knnx(road_coords, presence_coords, k = 1)
spiders$dist_to_road_km <- road_nn$nn.dist[,1] / 1000
cat(" done\n")
cat(" Finding nearest rivers...")
river_nn <- get.knnx(river_coords, presence_coords, k = 1)
spiders$dist_to_river_km <- river_nn$nn.dist[,1] / 1000
cat(" done\n")
# D5: Generate random points within study region
cat("\n5. Generating random points...\n")
n_random <- 1000
set.seed(456)
random_coords <- data.frame(
x = runif(n_random, xmin_buffer, xmax_buffer),
y = runif(n_random, ymin_buffer, ymax_buffer)
)
# D6: Calculate distances for random points
cat("\n6. Calculating distances for random points...\n")
random_matrix <- as.matrix(random_coords)
cat(" Finding nearest roads...")
random_road_nn <- get.knnx(road_coords, random_matrix, k = 1)
random_coords$dist_to_road_km <- random_road_nn$nn.dist[,1] / 1000
cat(" done\n")
cat(" Finding nearest rivers...")
random_river_nn <- get.knnx(river_coords, random_matrix, k = 1)
random_coords$dist_to_river_km <- random_river_nn$nn.dist[,1] / 1000
cat(" done\n")
random_coords$type <- "random"
# D7: Statistical tests
cat("\n7. Running statistical tests...\n")
spiders$type <- "observed"
ks_road <- ks.test(spiders$dist_to_road_km, random_coords$dist_to_road_km)
ks_river <- ks.test(spiders$dist_to_river_km, random_coords$dist_to_river_km)
cat("\nRESULTS:\n")
cat(sprintf(" Roads KS test: D = %.3f, p = %.4f %s\n",
ks_road$statistic, ks_road$p.value,
ifelse(ks_road$p.value < 0.05, "★ BIAS DETECTED", "")))
cat(sprintf(" Rivers KS test: D = %.3f, p = %.4f %s\n",
ks_river$statistic, ks_river$p.value,
ifelse(ks_river$p.value < 0.05, "★ BIAS DETECTED", "")))
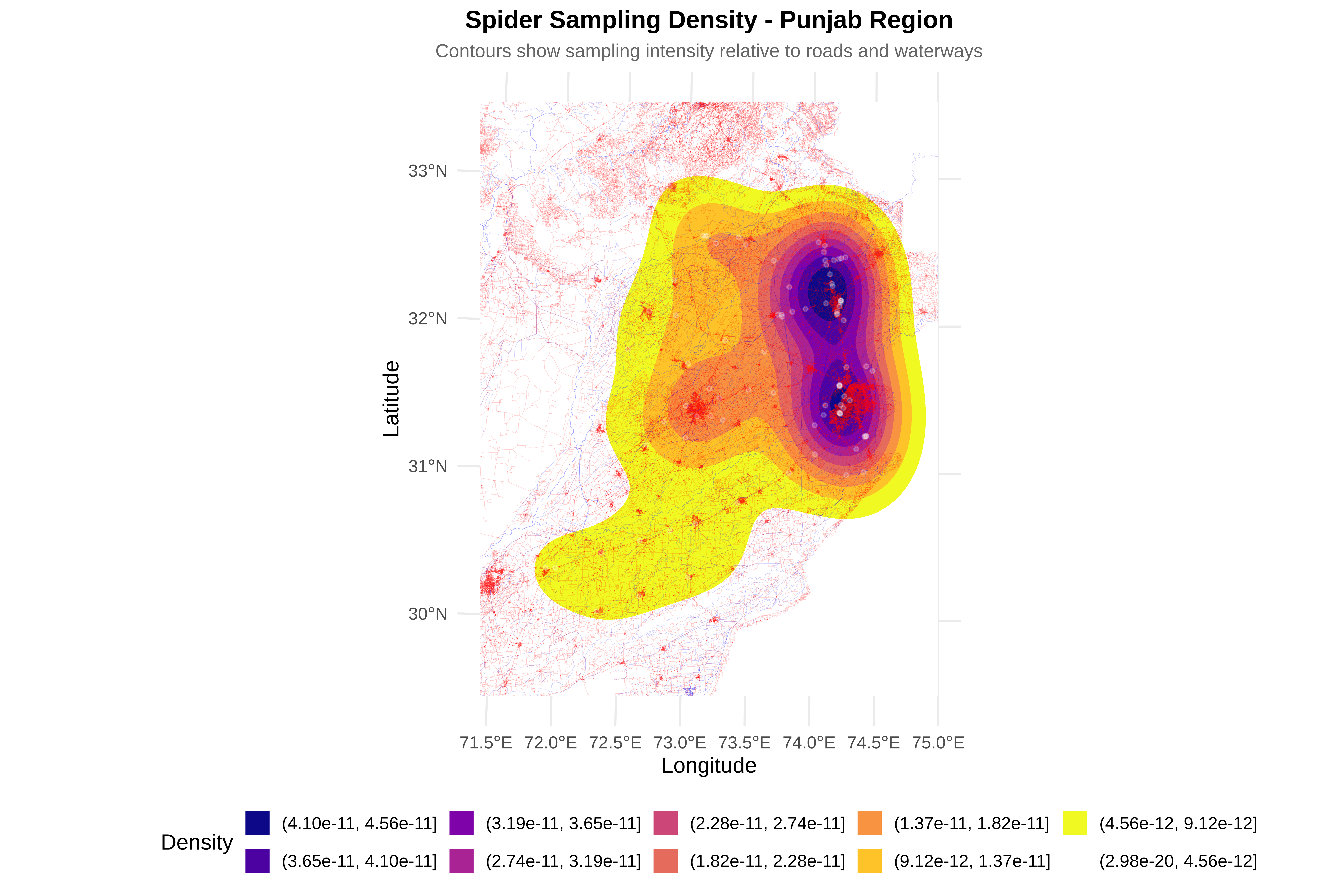
# D8: Visualize bias
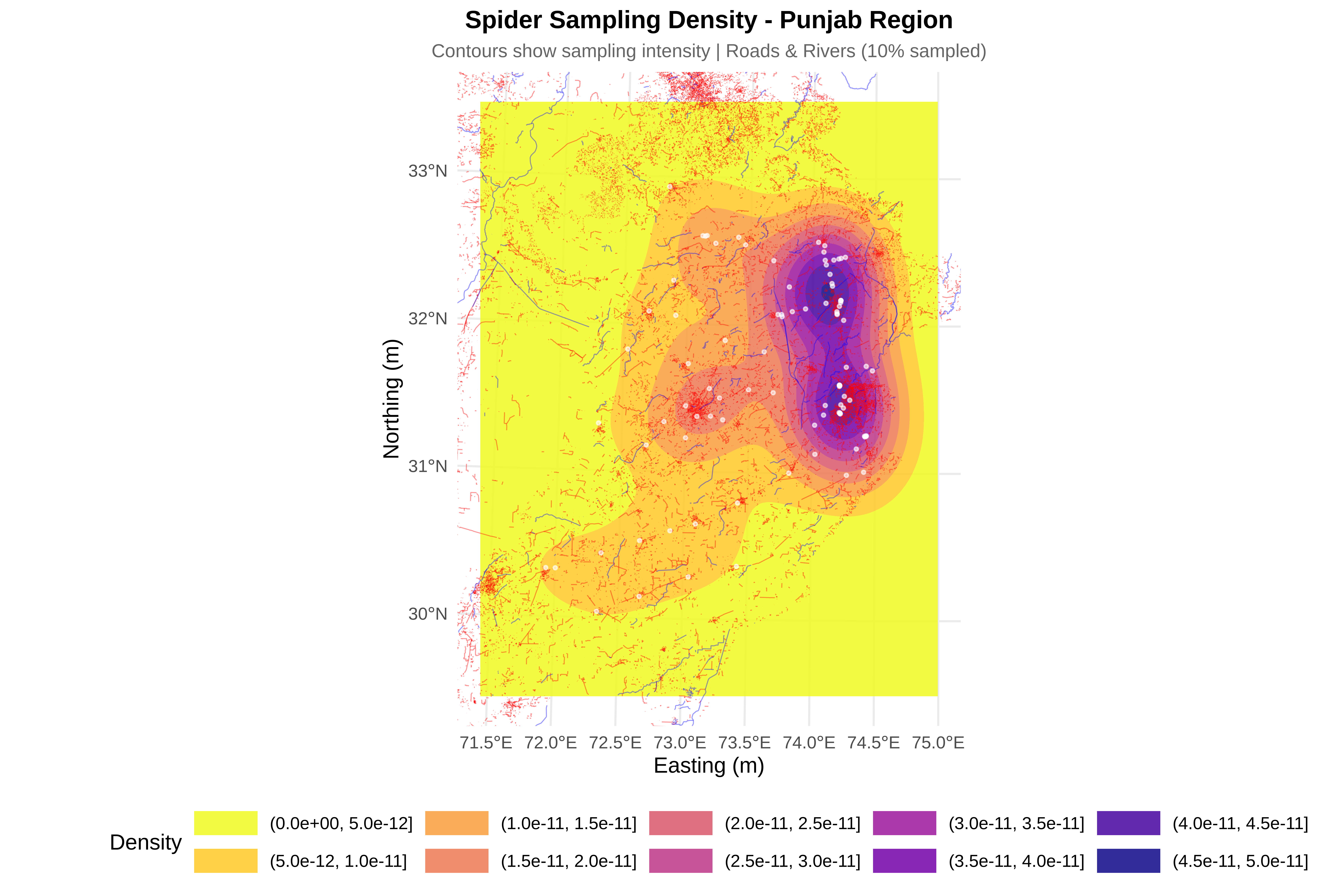
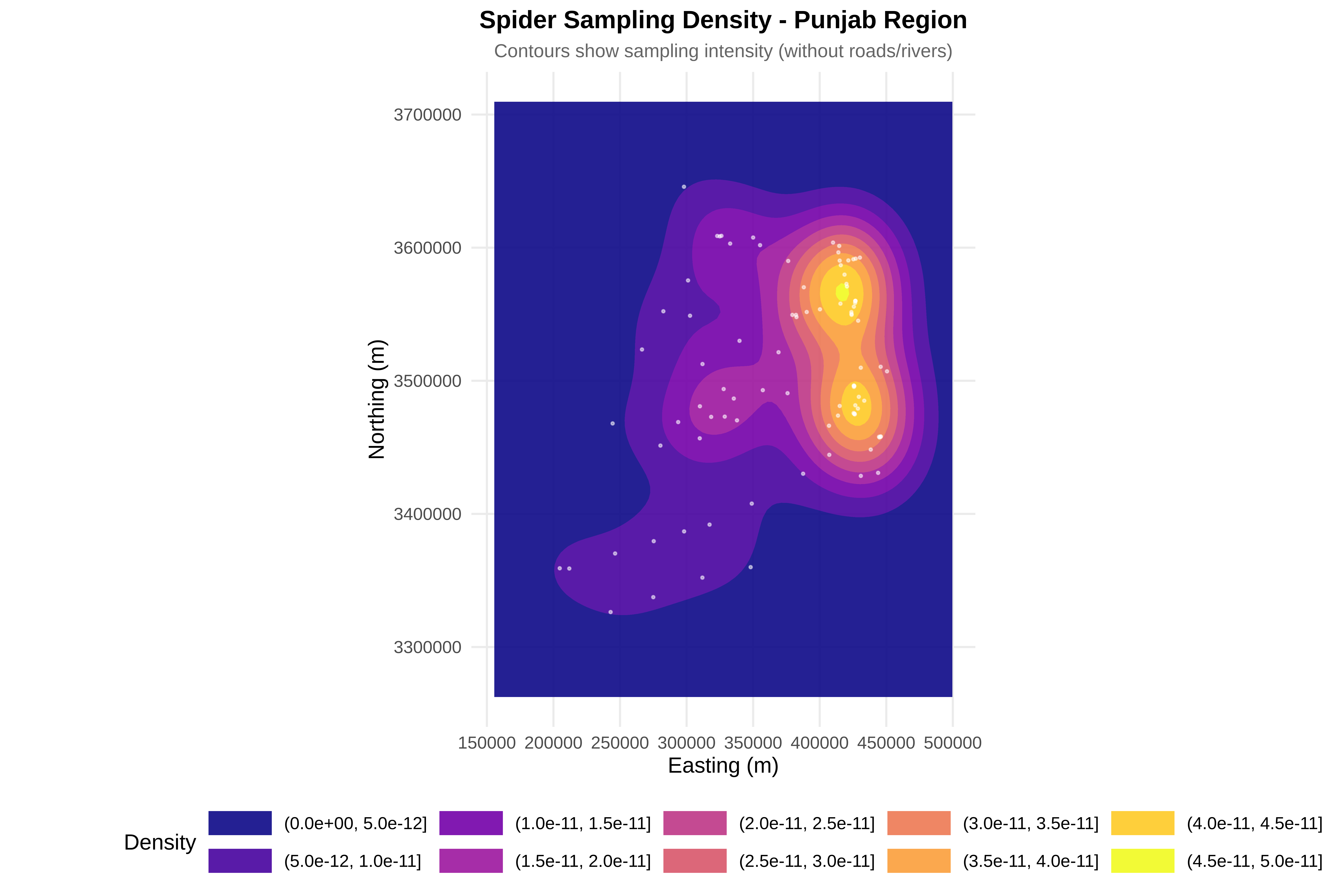
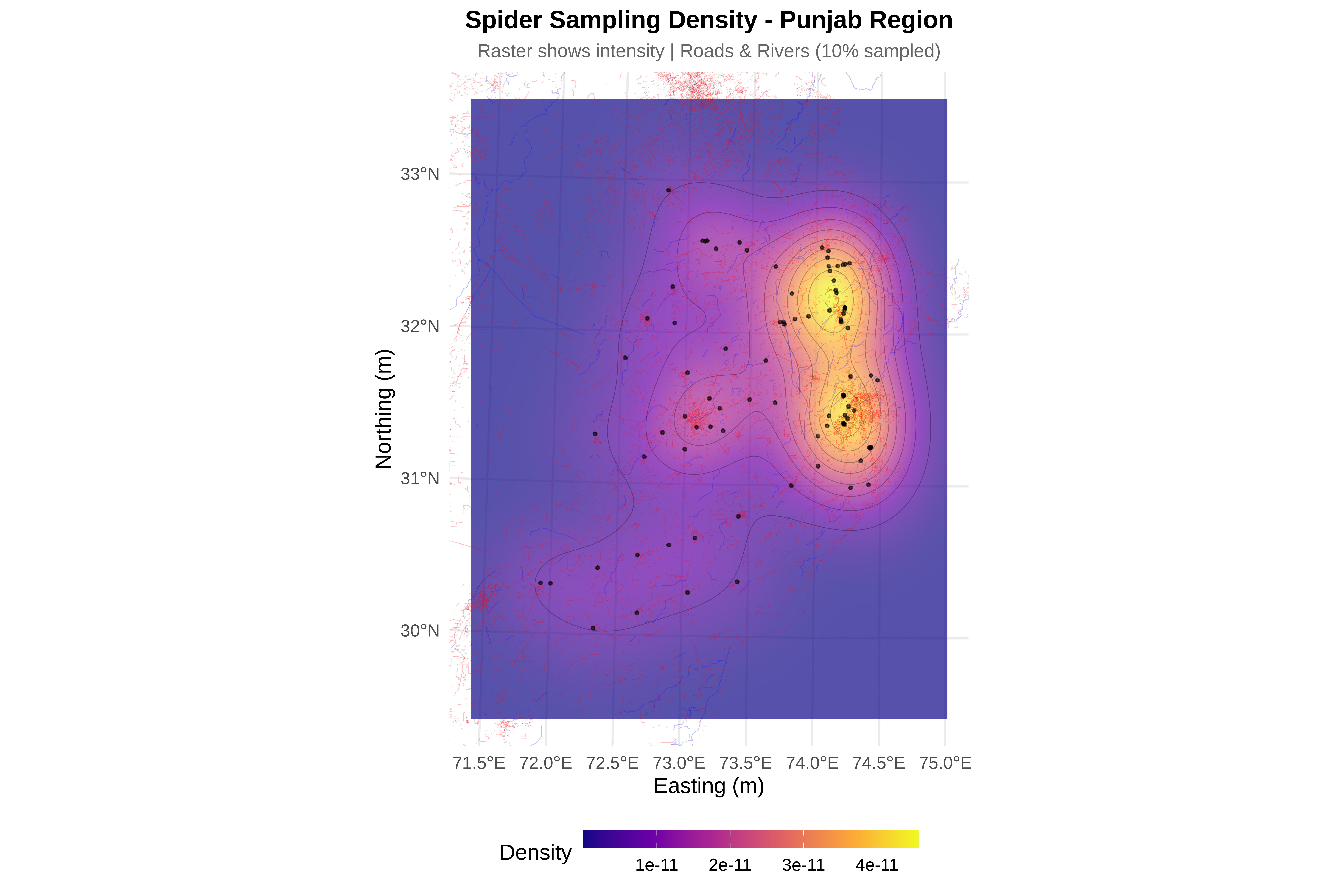
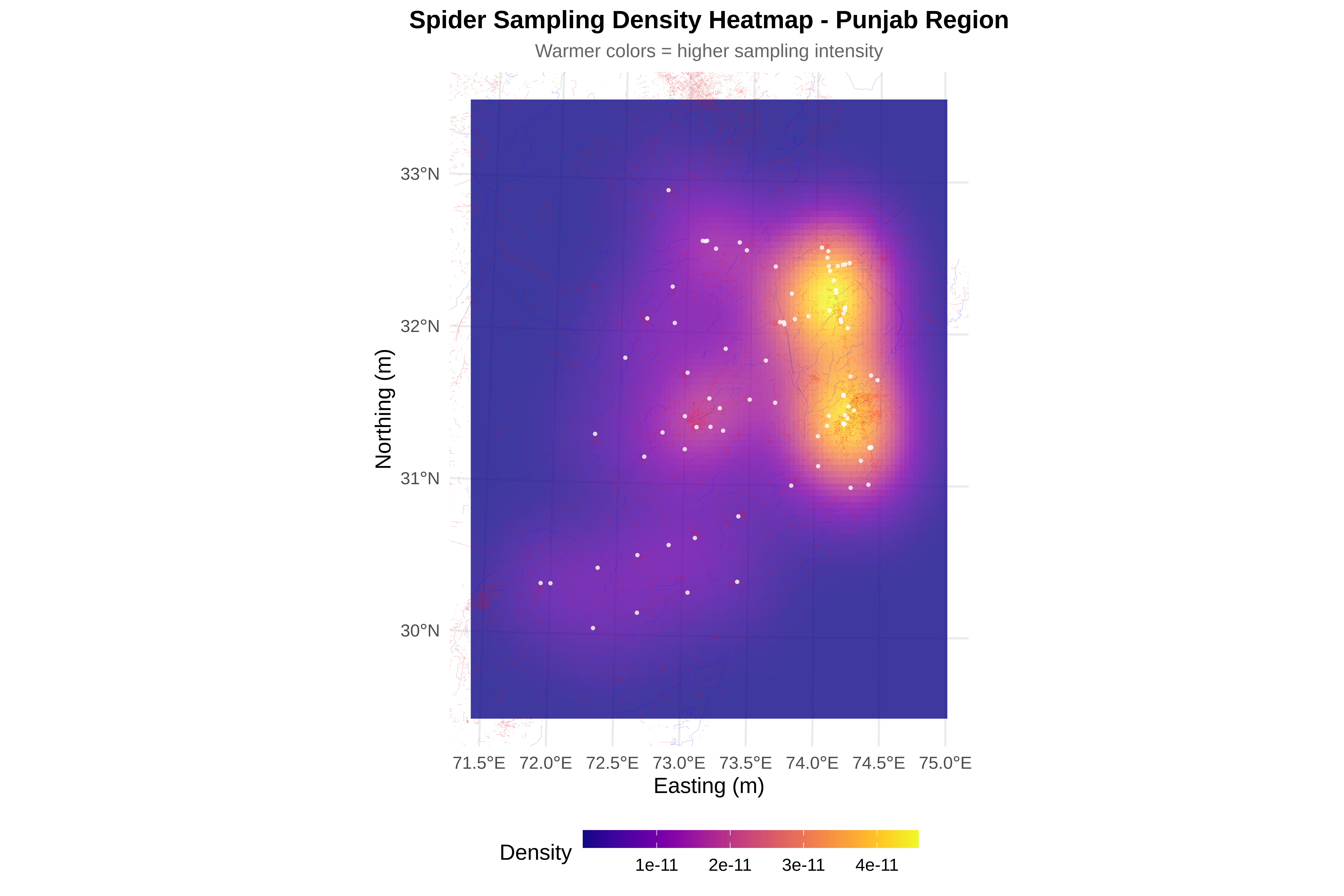
cat("\n8. Creating visualizations...\n")
bias_df <- rbind(
data.frame(dist = spiders$dist_to_road_km, type = spiders$type, feature = "Roads"),
data.frame(dist = random_coords$dist_to_road_km, type = random_coords$type, feature = "Roads"),
data.frame(dist = spiders$dist_to_river_km, type = spiders$type, feature = "Rivers"),
data.frame(dist = random_coords$dist_to_river_km, type = random_coords$type, feature = "Rivers")
)
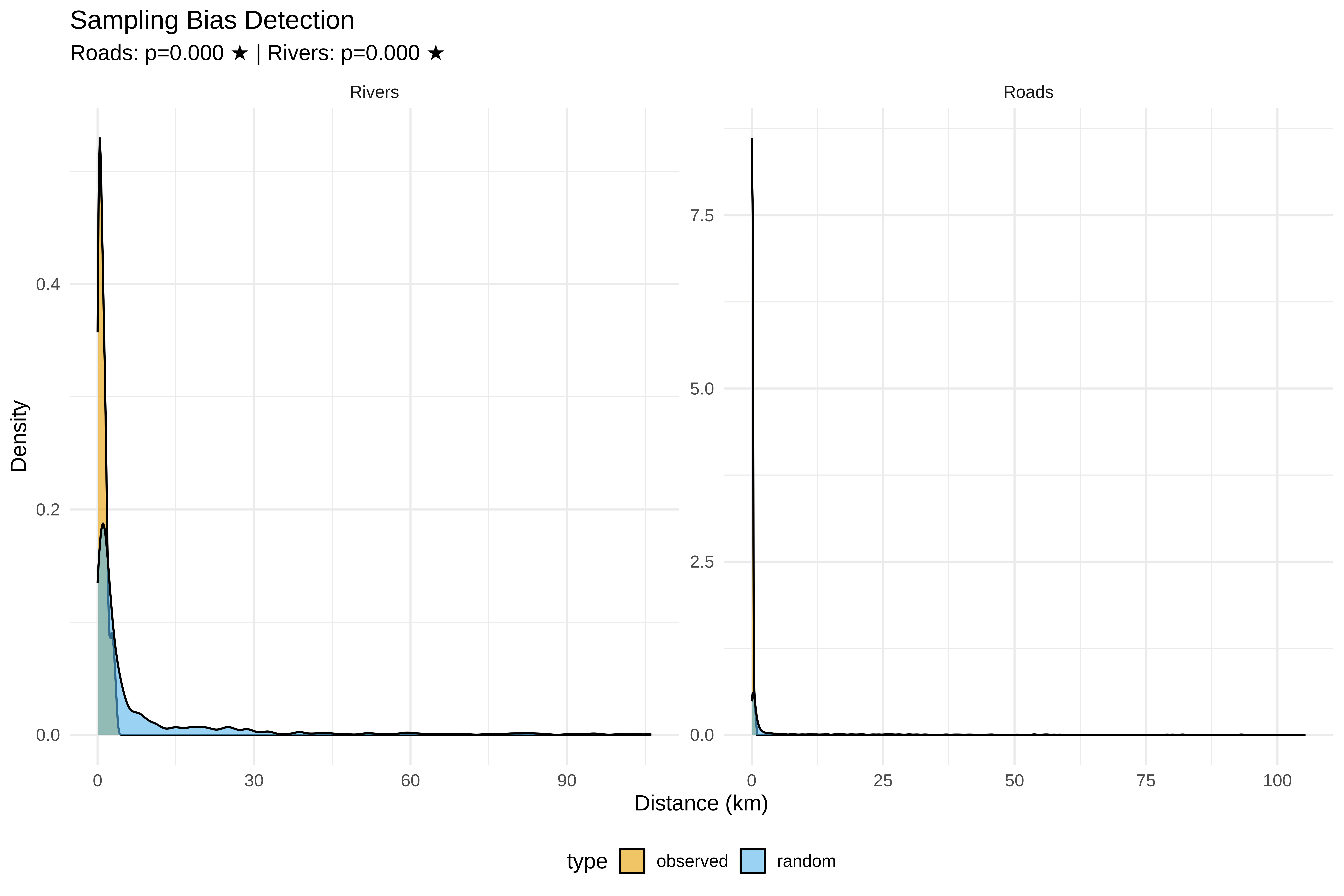
p_bias <- ggplot(bias_df, aes(x = dist, fill = type)) +
geom_density(alpha = 0.6) +
facet_wrap(~feature, scales = "free") +
scale_fill_manual(values = c("observed" = "#E69F00", "random" = "#56B4E9")) +
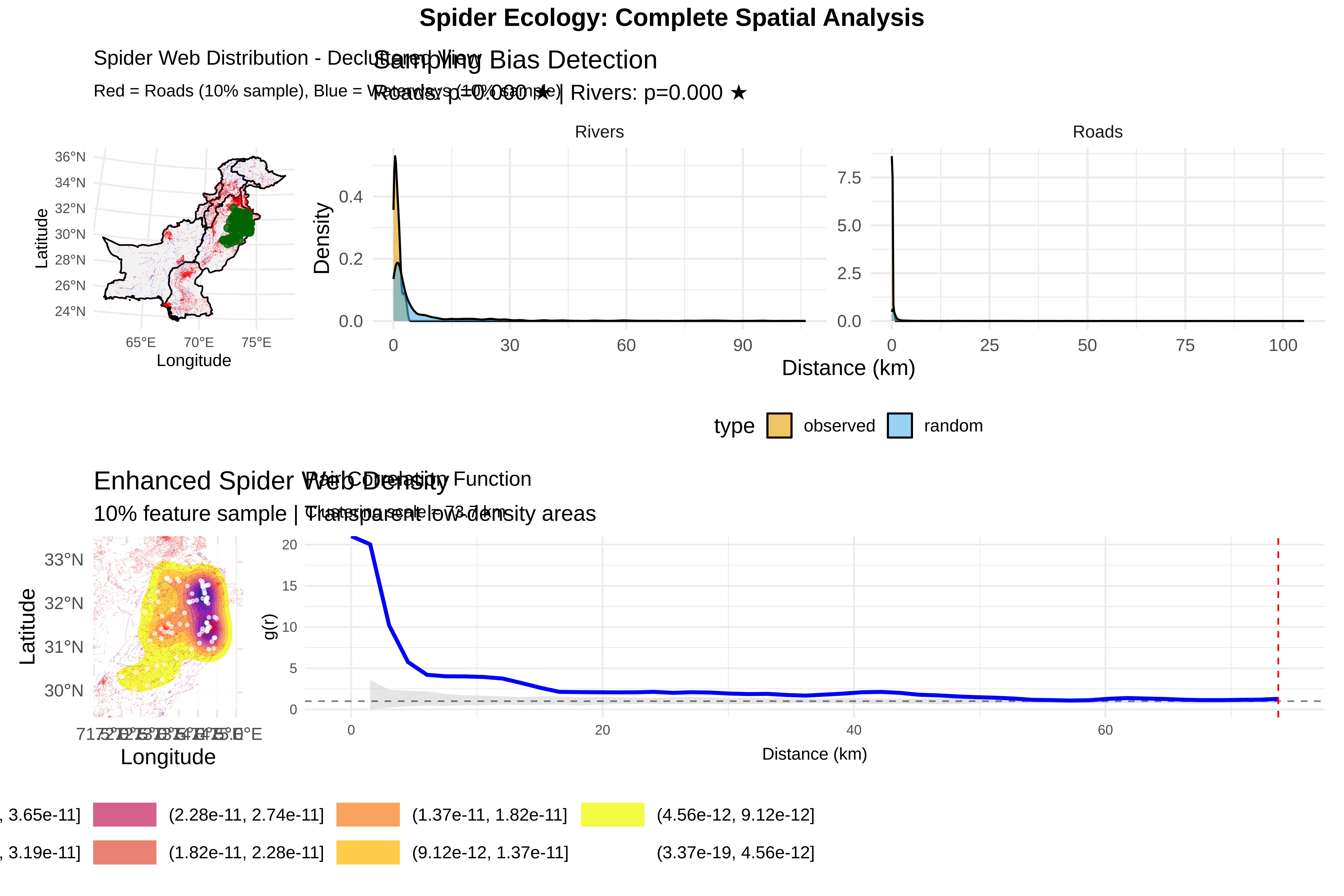
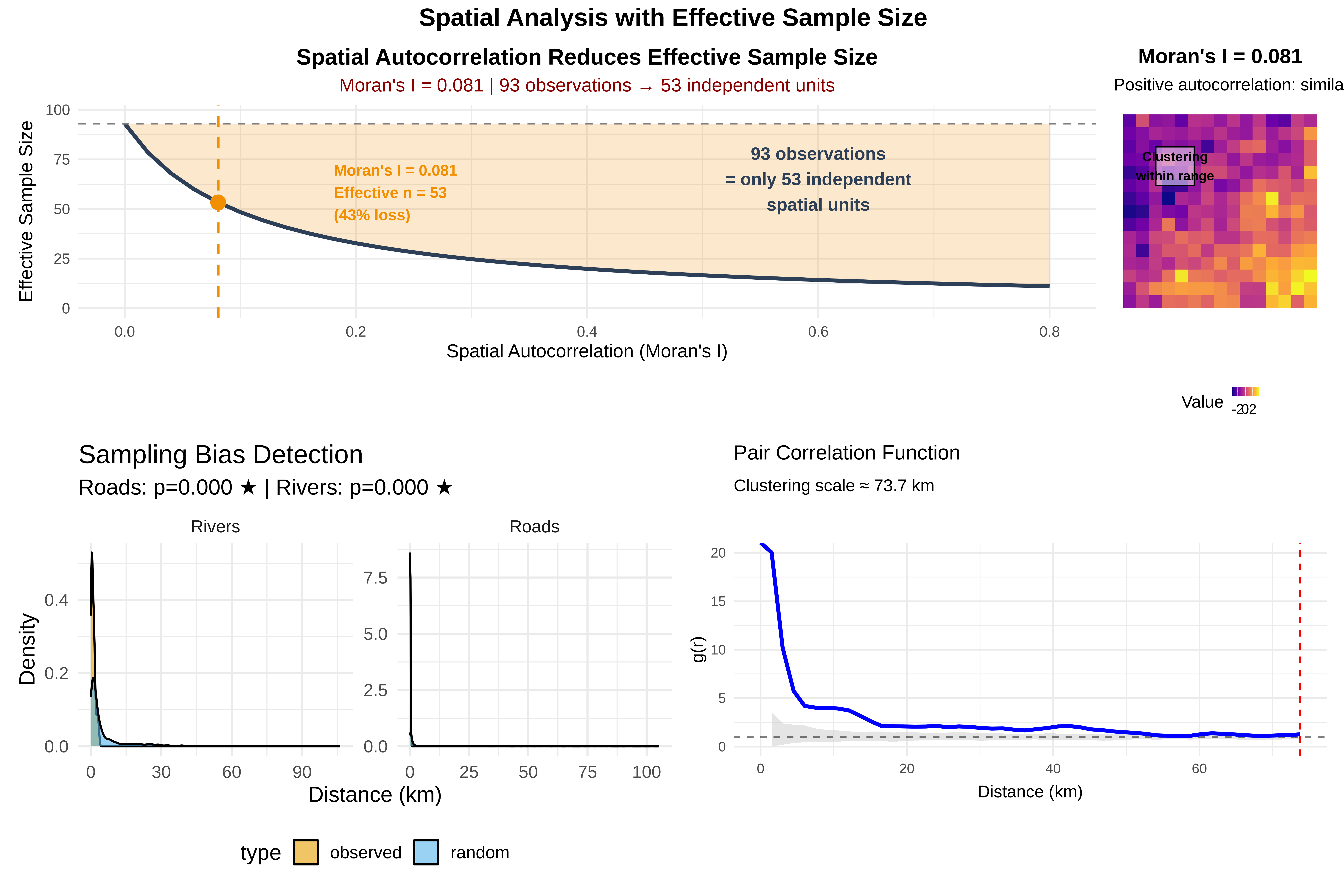
labs(title = "Sampling Bias Detection",
subtitle = sprintf("Roads: p=%.3f %s | Rivers: p=%.3f %s",
ks_road$p.value, ifelse(ks_road$p.value < 0.05, "★", ""),
ks_river$p.value, ifelse(ks_river$p.value < 0.05, "★", "")),
x = "Distance (km)", y = "Density") +
theme_minimal(base_size = 14) +
theme(legend.position = "bottom")
print(p_bias)