# ==============================================================================
# STEP 6: PERFORM PCA ON AVAILABLE ENVIRONMENT
# ==============================================================================
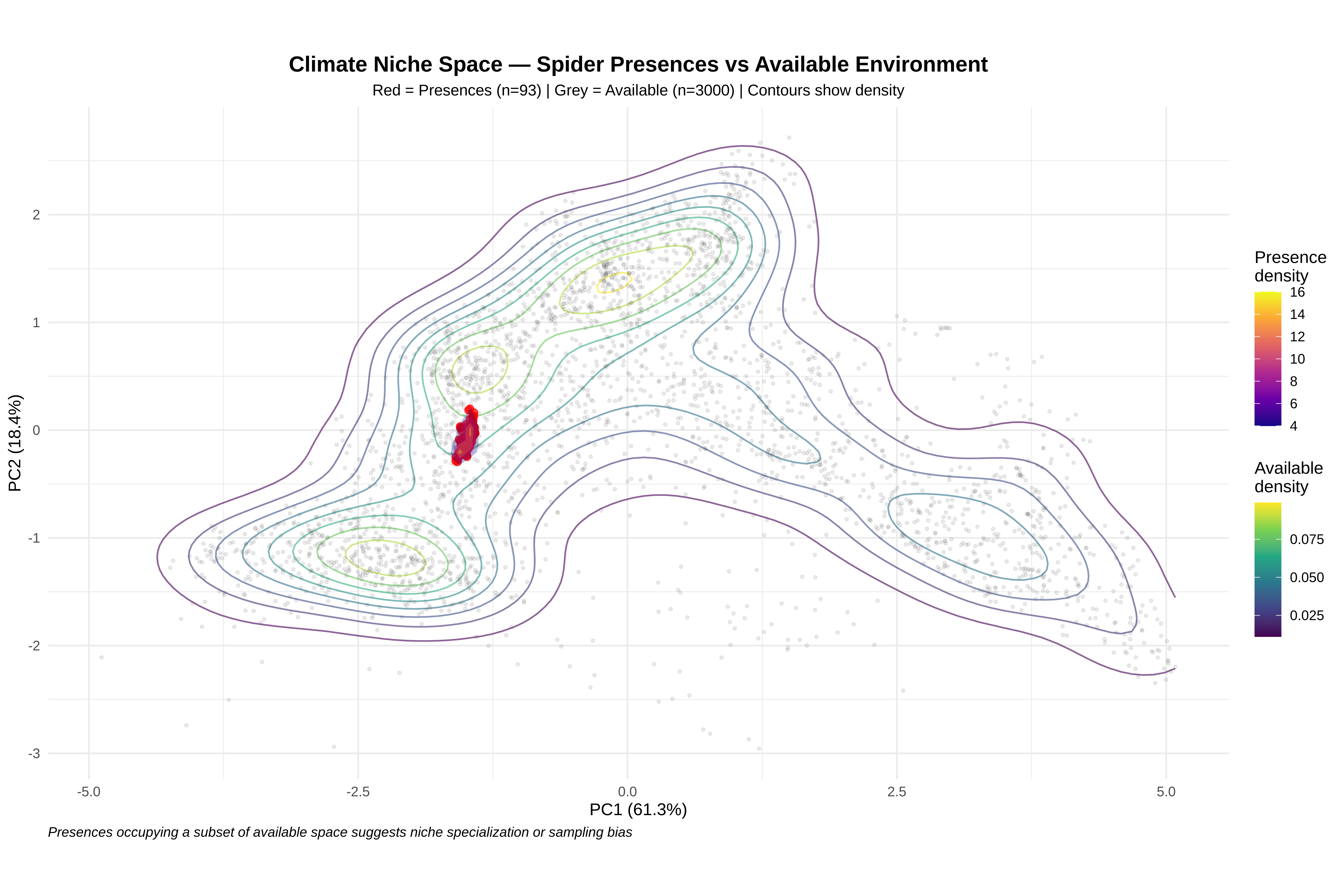
# We run PCA on the background (available) environment because:
# 1. It represents the FULL range of conditions in the study area
# 2. Presences are then PROJECTED into this space
# 3. This allows us to see WHERE presences sit relative to availability
# Scale variables to zero mean, unit variance (essential for PCA)
# Different units (°C, mm, m) would otherwise dominate based on magnitude
avail_scaled <- scale(avail_env)
# Store scaling parameters — we'll need these to project presences
scale_center <- attr(avail_scaled, "scaled:center")
scale_sd <- attr(avail_scaled, "scaled:scale")
# Perform PCA
pca <- prcomp(avail_scaled)
# Calculate variance explained by each PC
var_exp <- round(summary(pca)$importance[2, ] * 100, 1)
cum_var <- round(summary(pca)$importance[3, ] * 100, 1)
cat("\n", paste(rep("=", 60), collapse = ""), "\n")
cat("PCA VARIANCE EXPLAINED\n")
cat(paste(rep("=", 60), collapse = ""), "\n\n")
cat("Individual variance explained:\n")
for(i in 1:length(var_exp)) {
cat(sprintf(" PC%d: %5.1f%% (cumulative: %5.1f%%)\n", i, var_exp[i], cum_var[i]))
}
cat(sprintf("\nFirst 2 PCs explain: %.1f%% of total variance\n", cum_var[2]))
cat(sprintf("First 3 PCs explain: %.1f%% of total variance\n", cum_var[3]))
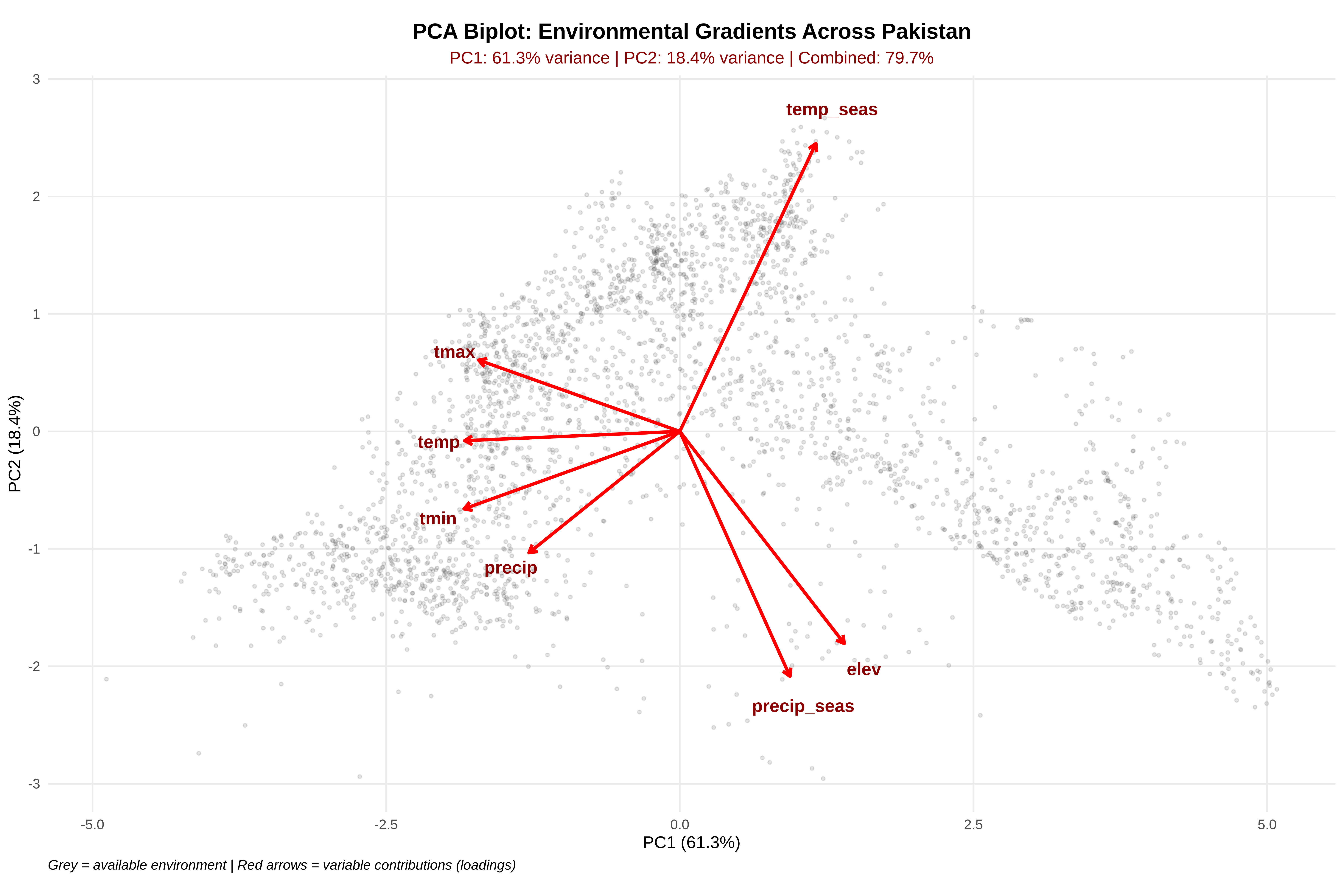
# --- Create PCA Biplot ---
# Extract scores (point positions in PC space) and loadings (variable arrows)
scores <- as.data.frame(pca$x[, 1:2])
load <- as.data.frame(pca$rotation[, 1:2])
load$var <- rownames(load)
# Determine scaling factor for arrows (so they're visible relative to points)
scale_factor <- min(
(max(scores$PC1) - min(scores$PC1)) / (max(load$PC1) - min(load$PC1)),
(max(scores$PC2) - min(scores$PC2)) / (max(load$PC2) - min(load$PC2))
) * 0.8
# Build biplot
p_biplot <- ggplot() +
# Background points (available environment)
geom_point(data = scores, aes(x = PC1, y = PC2),
alpha = 0.15, size = 0.8, color = "gray30") +
# Origin marker
geom_point(x = 0, y = 0, size = 2, color = "red") +
# Variable loading arrows
geom_segment(data = load,
aes(x = 0, y = 0,
xend = PC1 * scale_factor,
yend = PC2 * scale_factor),
arrow = arrow(length = unit(0.2, "cm")),
color = "red", linewidth = 1) +
# Variable labels
geom_text(data = load,
aes(x = PC1 * scale_factor * 1.12,
y = PC2 * scale_factor * 1.12,
label = var),
color = "darkred", size = 4, fontface = "bold") +
# Axis labels with variance explained
labs(
title = "PCA Biplot: Environmental Gradients Across Pakistan",
subtitle = paste0("PC1: ", var_exp[1], "% variance | PC2: ",
var_exp[2], "% variance | Combined: ",
cum_var[2], "%"),
x = paste0("PC1 (", var_exp[1], "%)"),
y = paste0("PC2 (", var_exp[2], "%)"),
caption = "Grey = available environment | Red arrows = variable contributions (loadings)"
) +
# Equal aspect ratio is essential for interpreting angles between arrows
coord_fixed() +
theme_minimal() +
theme(
plot.title = element_text(hjust = 0.5, face = "bold", size = 14),
plot.subtitle = element_text(hjust = 0.5, color = "darkred", size = 11),
plot.caption = element_text(hjust = 0, face = "italic"),
panel.grid.minor = element_blank()
)
print(p_biplot)