# ==============================================================================
# STEP 14: RUN SPATIAL CROSS-VALIDATION ACROSS ALL MODELS
# ==============================================================================
# Helper: calculate AUC
calc_auc <- function(y, p) {
tryCatch({
as.numeric(pROC::auc(pROC::roc(y, p, quiet = TRUE)))
}, error = function(e) NA_real_)
}
# Helper: fit all models and return predictions for test set
fit_and_predict <- function(train, test, pred_vars) {
out <- list()
# Ensure pa is factor for RF
train$pa <- as.factor(train$pa)
# 1. GLM (quadratic terms)
glm_f <- as.formula(paste("pa ~",
paste(c(pred_vars, paste0("I(", pred_vars, "^2)")),
collapse = " + ")))
tryCatch({
m_glm <- glm(glm_f, data = train, family = binomial)
out$GLM <- predict(m_glm, newdata = test, type = "response")
}, error = function(e) { out$GLM <- rep(NA, nrow(test)) })
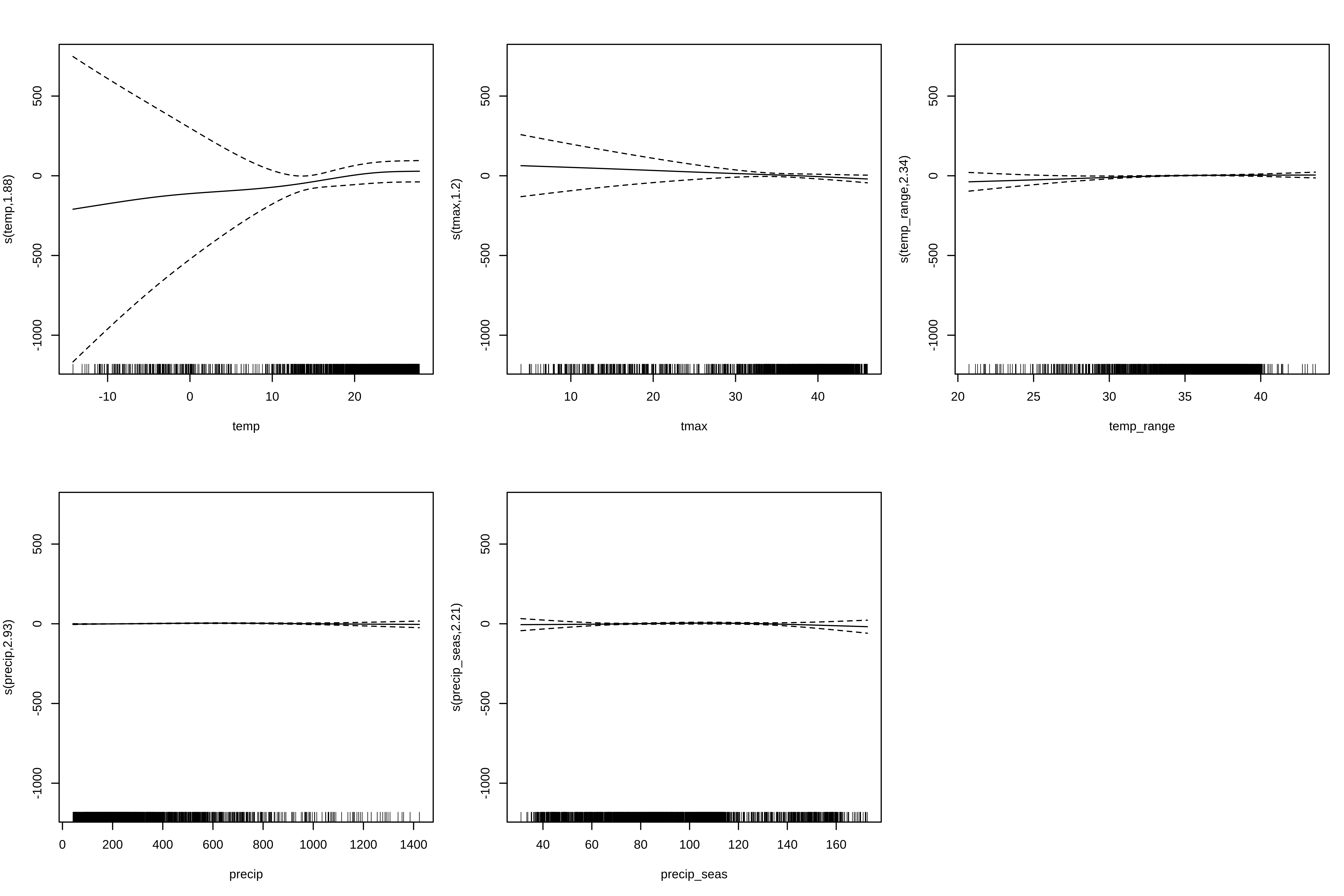
# 2. GAM (smooth terms)
gam_terms <- paste0("s(", pred_vars, ", k = 4)")
gam_f <- as.formula(paste("pa ~", paste(gam_terms, collapse = " + ")))
tryCatch({
m_gam <- mgcv::gam(gam_f, data = train, family = binomial, method = "REML")
out$GAM <- predict(m_gam, newdata = test, type = "response")
}, error = function(e) { out$GAM <- rep(NA, nrow(test)) })
# 3. Random Forest
tryCatch({
m_rf <- ranger::ranger(
as.factor(pa) ~ .,
data = train[, c("pa", pred_vars)],
probability = TRUE, num.trees = 400
)
out$RF <- predict(m_rf, data = test)$predictions[, "1"]
}, error = function(e) { out$RF <- rep(NA, nrow(test)) })
# 4. Maxnet
tryCatch({
f <- maxnet::maxnet.formula(train$pa, train[, pred_vars], classes = "lqph")
m_mx <- maxnet::maxnet(p = train$pa, data = train[, pred_vars],
f = f, regmult = 1.5)
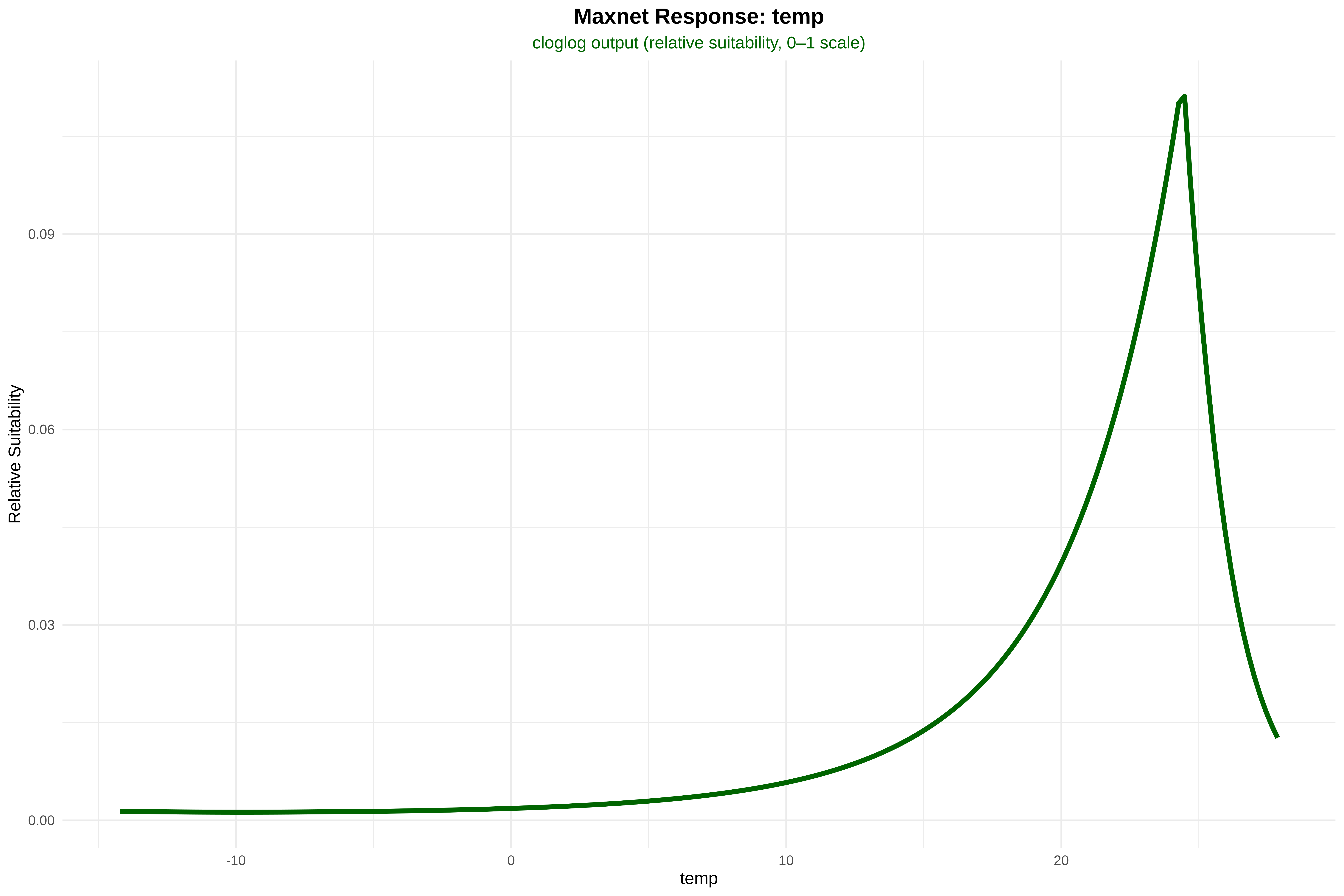
out$Maxnet <- as.numeric(predict(m_mx, test[, pred_vars], type = "cloglog"))
}, error = function(e) { out$Maxnet <- rep(NA, nrow(test)) })
return(out)
}
# --- Run cross-validation ---
cat("\nRunning spatial cross-validation...\n")
cv_results <- data.frame()
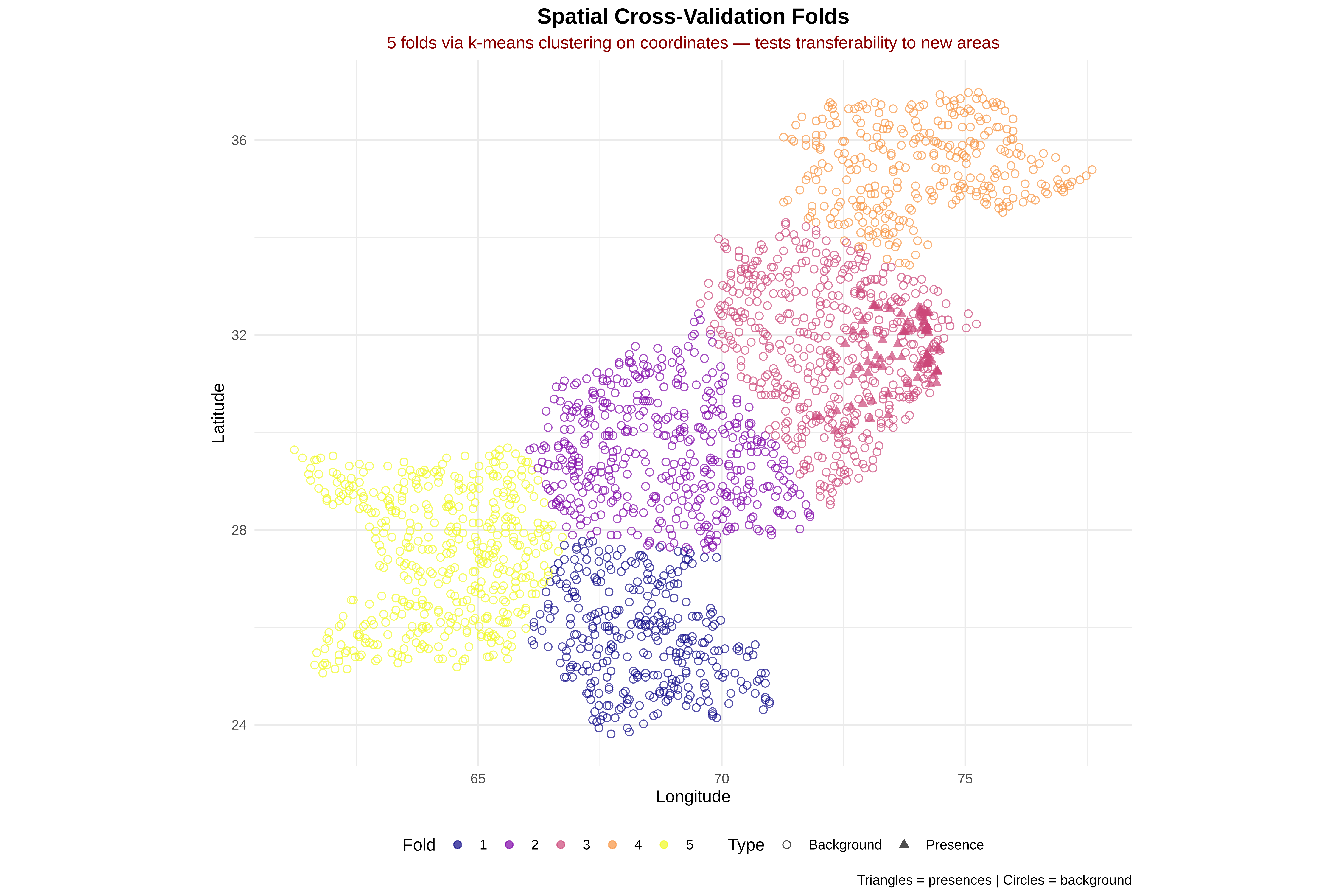
for(k in sort(unique(dat$fold))) {
cat(" Fold", k, "...")
train <- dat %>% filter(fold != k)
test <- dat %>% filter(fold == k)
preds <- fit_and_predict(train, test, pred_vars)
for(model_name in names(preds)) {
auc_val <- calc_auc(test$pa, preds[[model_name]])
cv_results <- rbind(cv_results, data.frame(
fold = k, model = model_name, AUC = auc_val
))
cat(paste0(" ", model_name, "=", round(auc_val, 3)))
}
cat("\n")
}
# --- Summarize results ---
results_summary <- cv_results %>%
group_by(model) %>%
summarise(
mean_AUC = mean(AUC, na.rm = TRUE),
sd_AUC = sd(AUC, na.rm = TRUE),
n_folds = sum(!is.na(AUC)),
.groups = "drop"
) %>%
arrange(desc(mean_AUC))
cat("\n", paste(rep("=", 60), collapse = ""), "\n")
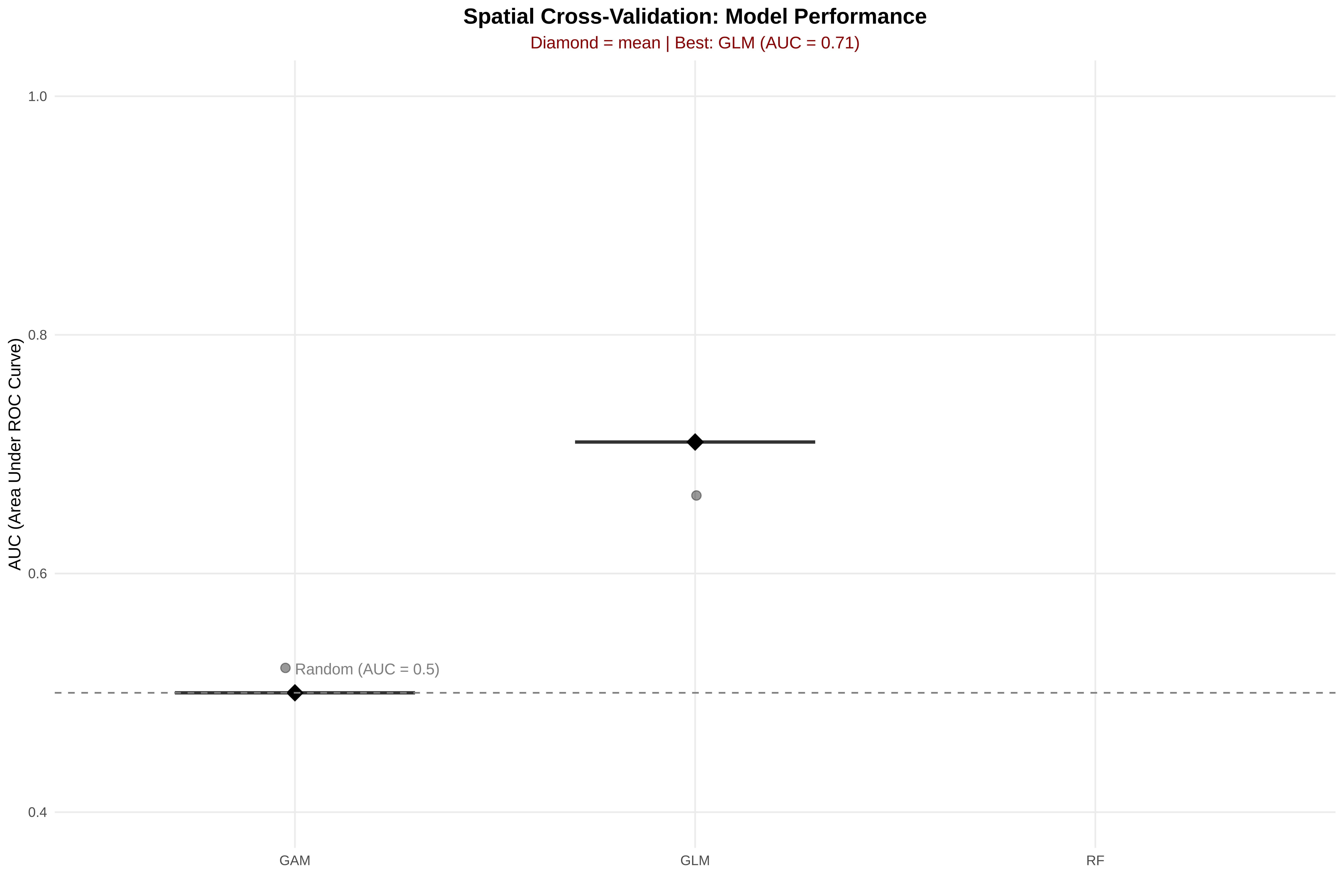
cat("SPATIAL CV RESULTS — MODEL COMPARISON\n")
cat(paste(rep("=", 60), collapse = ""), "\n\n")
print(as.data.frame(results_summary))
best_model_name <- results_summary$model[1]
cat("\n✓ Best model:", best_model_name,
"(AUC =", round(results_summary$mean_AUC[1], 3),
"±", round(results_summary$sd_AUC[1], 3), ")\n")
# ==============================================================================
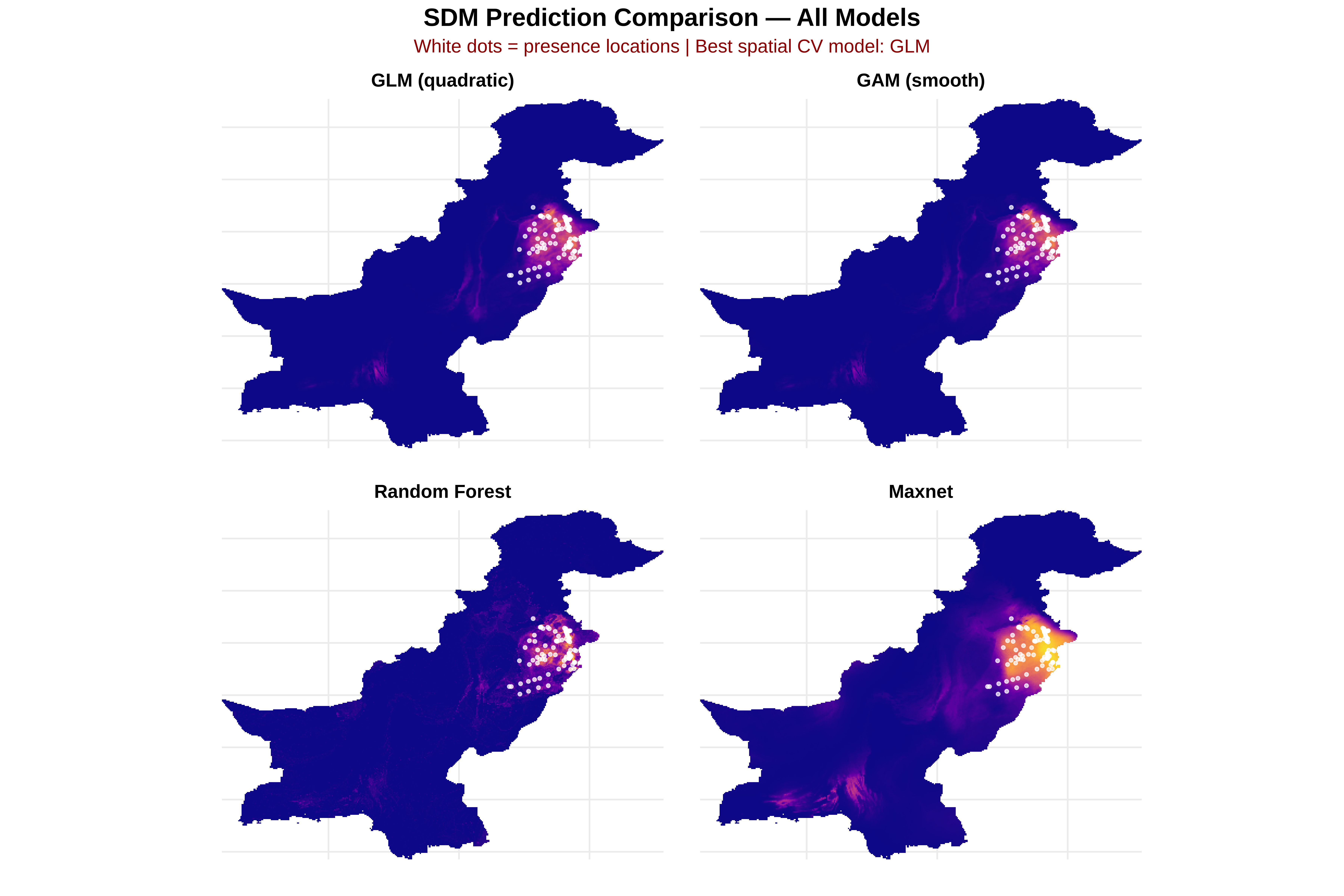
# STEP 14B: FIT FINAL MODELS ON FULL DATA FOR PREDICTION
# ==============================================================================
# --- Quadratic GLM formula ---
glm_quad_formula <- as.formula(paste("pa ~",
paste(c(pred_vars, paste0("I(", pred_vars, "^2)")), collapse = " + ")))
# --- GAM formula ---
gam_terms <- paste0("s(", pred_vars, ", k = 4)")
gam_formula <- as.formula(paste("pa ~", paste(gam_terms, collapse = " + ")))
# 1. GLM (quadratic)
glm_model <- glm(glm_quad_formula, data = dat, family = binomial)
# 2. GAM
gam_model <- mgcv::gam(gam_formula, data = dat, family = binomial, method = "REML")
# 3. Random Forest
rf_model <- ranger::ranger(
as.factor(pa) ~ .,
data = dat[, c("pa", pred_vars)],
probability = TRUE, num.trees = 400
)
# 4. Maxnet
f <- maxnet::maxnet.formula(dat$pa, dat[, pred_vars], classes = "lqph")
maxnet_model <- maxnet::maxnet(p = dat$pa, data = dat[, pred_vars], f = f, regmult = 1.5)
cat("✓ Final models fitted on full dataset for prediction.\n")
# print(cat("✓ Final models fitted on full dataset for prediction.\n"))